Soumick Chatterjee, Hendrik Mattern, Marc Dörner, Alessandro Sciarra, Florian Dubost, Hannes Schnurre, Rupali Khatun, Chun-Chih Yu, Tsung-Lin Hsieh, Yi-Shan Tsai, Yi-Zeng Fang, Yung-Ching Yang, Juinn-Dar Huang, Marshall Xu, Siyu Liu, Fernanda L Ribeiro, Saskia Bollmann, Karthikesh Varma Chintalapati, Chethan Mysuru Radhakrishna, Sri Chandana Hudukula Ram Kumara, Raviteja Sutrave, Abdul Qayyum, Moona Mazher, Imran Razzak, Cristobal Rodero, Steven Niederren, Fengming Lin, Yan Xia, Jiacheng Wang, Riyu Qiu, Liansheng Wang, Arya Yazdan Panah, Rosana El Jurdi, Guanghui Fu, Janan Arslan, Ghislain Vaillant, Romain Valabregue, Didier Dormont, Bruno Stankoff, Olivier Colliot, Luisa Vargas, Isai Daniel Chacón, Ioannis Pitsiorlas, Pablo Arbeláez, Maria A Zuluaga, Stefanie Schreiber, Oliver Speck, Andreas Nürnberger
arXiv (2024)
Abstract
The human brain receives nutrients and oxygen through an intricate network of blood vessels. Pathology affecting small vessels, at
the mesoscopic scale, represents a critical vulnerability within the cerebral blood supply and can lead to severe conditions, such as
Cerebral Small Vessel Diseases. The advent of 7 Tesla MRI systems has enabled the acquisition of higher spatial resolution images,
making it possible to visualise such vessels in the brain. However, the lack of publicly available annotated datasets has impeded
the development of robust, machine learning-driven segmentation algorithms. To address the complexities of mesoscopic vessel
segmentation and to highlight the need for advanced techniques to manage the high noise levels and poor vessel-to-background
contrast inherent in ”ultra-high-resolution” data, the SMILE-UHURA challenge was organised. This challenge, held in conjunction
with the ISBI 2023, in Cartagena de Indias, Colombia, aimed to provide a platform for researchers working on related topics.
The SMILE-UHURA challenge addresses the gap in publicly available annotated datasets by providing an annotated dataset of
Time-of-Flight angiography acquired with 7T MRI. This dataset was created through a combination of automated pre-segmentation
and extensive manual refinement. In this manuscript, sixteen submitted methods and two baseline methods are compared both
quantitatively and qualitatively on two different datasets: held-out test MRAs from the same dataset as the training data (with labels
kept secret) and a separate 7T ToF MRA dataset where both input volumes and labels are kept secret. The results demonstrate that
most of the submitted deep learning methods, trained on the provided training dataset, achieved reliable segmentation performance.
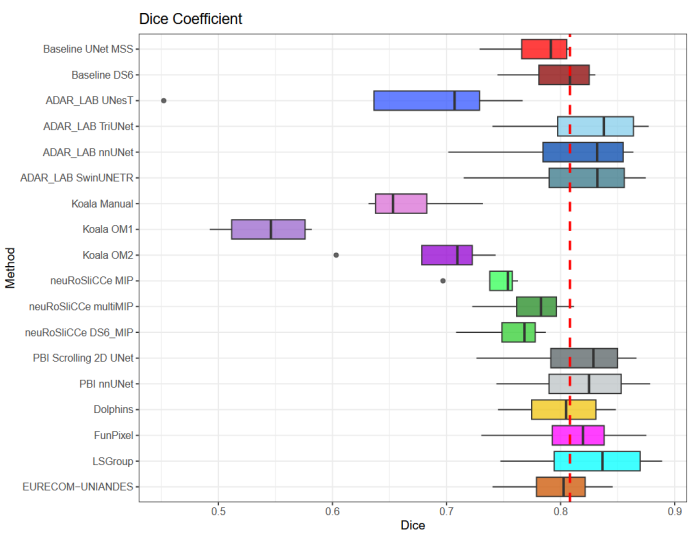
Dice scores reached up to 0.838 ± 0.066 and 0.716 ± 0.125 on the respective datasets, with an average performance of up to 0.804
± 0.15.